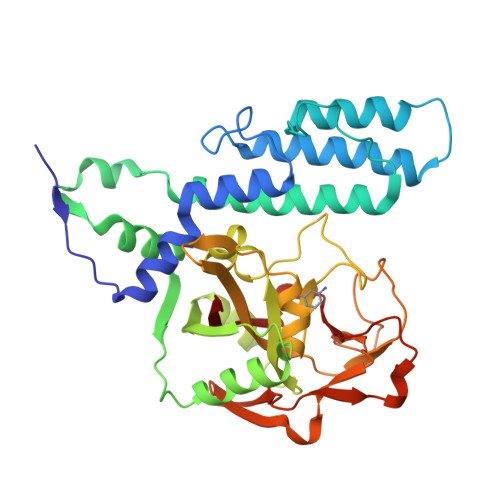
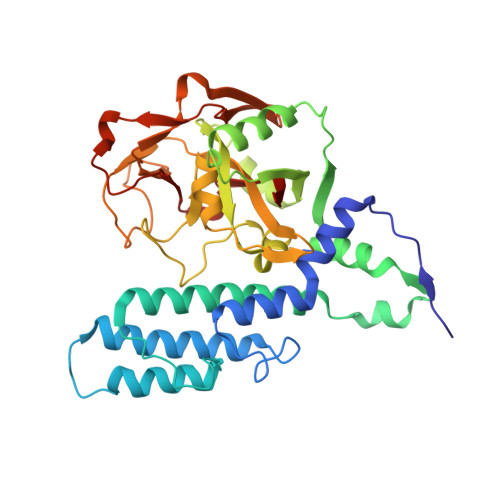
Structural basis for inhibitor specificity in human poly(ADP-ribose) polymerase-3.
Lehtio, L., Jemth, A.S., Collins, R., Loseva, O., Johansson, A., Markova, N., Hammarstrom, M., Flores, A., Holmberg-Schiavone, L., Weigelt, J., Helleday, T., Schuler, H., Karlberg, T.(2009) J Med Chem 52: 3108-3111
- PubMed: 19354255
- DOI: https://doi.org/10.1021/jm900052j
- Primary Citation of Related Structures:
3C49, 3C4H, 3CE0, 3FHB - PubMed Abstract:
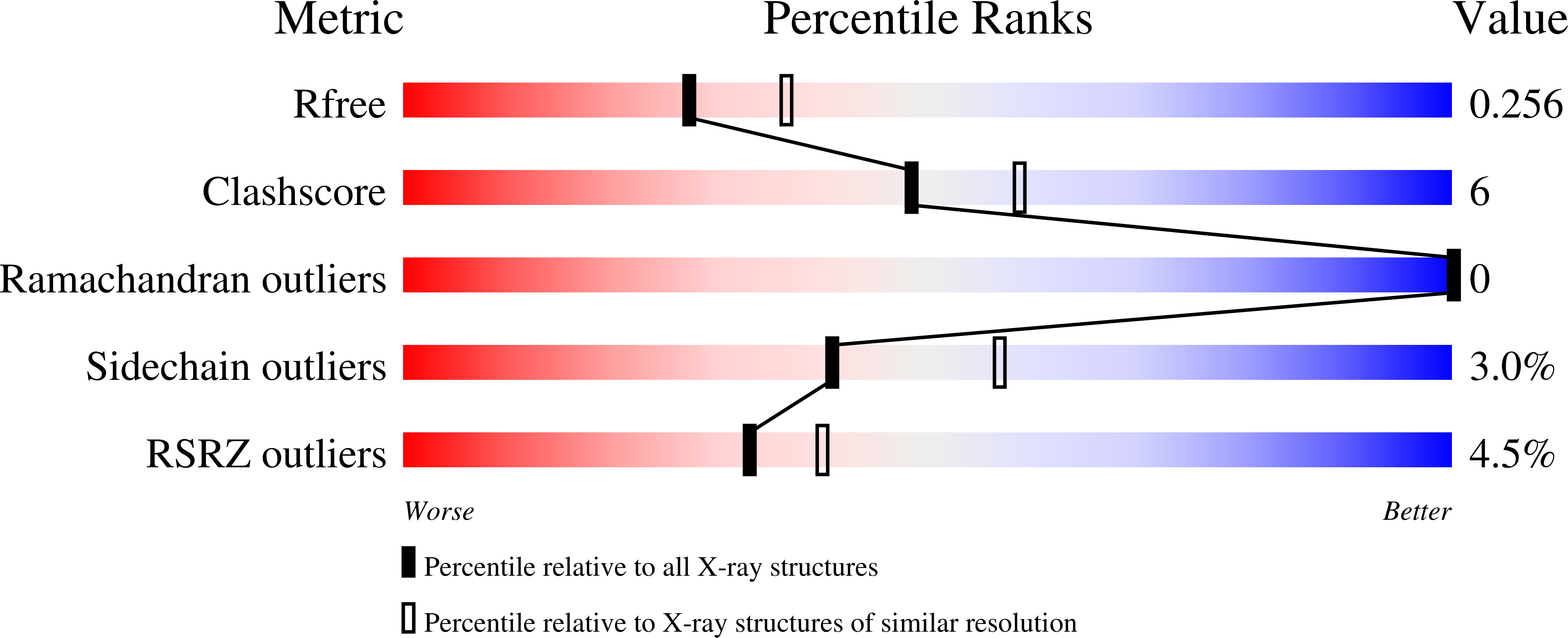
Poly(ADP-ribose) polymerases (PARPs) activate DNA repair mechanisms upon stress- and cytotoxin-induced DNA damage, and inhibition of PARP activity is a lead in cancer drug therapy. We present a structural and functional analysis of the PARP domain of human PARP-3 in complex with several inhibitors. Of these, KU0058948 is the strongest inhibitor of PARP-3 activity. The presented crystal structures highlight key features for potent inhibitor binding and suggest routes for creating isoenzyme-specific PARP inhibitors.
Organizational Affiliation:
Structural Genomics Consortium, Department of Medical Biochemistry and Biophysics, Karolinska Institutet, SE-17177 Stockholm, Sweden.